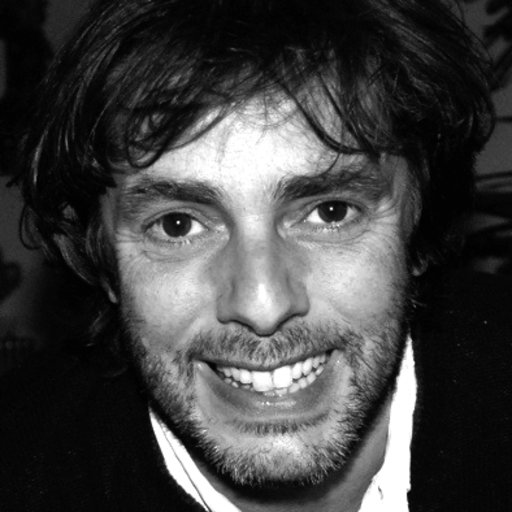

 Adriano Henriques and Mónica Serrano are long-term collaborators who warmly welcomed me to the Bacterial Development lab, where they are guiding my attempt to understand the origin, biology and ecology of bacterial spores. Our common understanding is that while I immerse myself in the thinking of the lab, I am available to support anyone needing advice on statistics and quantitative data analysis.
Adriano Henriques and Mónica Serrano are long-term collaborators who warmly welcomed me to the Bacterial Development lab, where they are guiding my attempt to understand the origin, biology and ecology of bacterial spores. Our common understanding is that while I immerse myself in the thinking of the lab, I am available to support anyone needing advice on statistics and quantitative data analysis.
 Alberto Darszon has been both a collaborator and a friend for several decades. We’ve spent sabbaticals in each other’s labs, building on a collaboration that began at a workshop in Cuernavaca, where I was mesmerised by his presentation of live imaging of sea urchin spermatozoa swimming in circles at hundreds of frames per second. Since then, we’ve worked together on modelling various aspects of sea urchin and human sperm physiology, as well as on imaging data analysis, contributing to the training of several PhD students.
Alberto Darszon has been both a collaborator and a friend for several decades. We’ve spent sabbaticals in each other’s labs, building on a collaboration that began at a workshop in Cuernavaca, where I was mesmerised by his presentation of live imaging of sea urchin spermatozoa swimming in circles at hundreds of frames per second. Since then, we’ve worked together on modelling various aspects of sea urchin and human sperm physiology, as well as on imaging data analysis, contributing to the training of several PhD students.
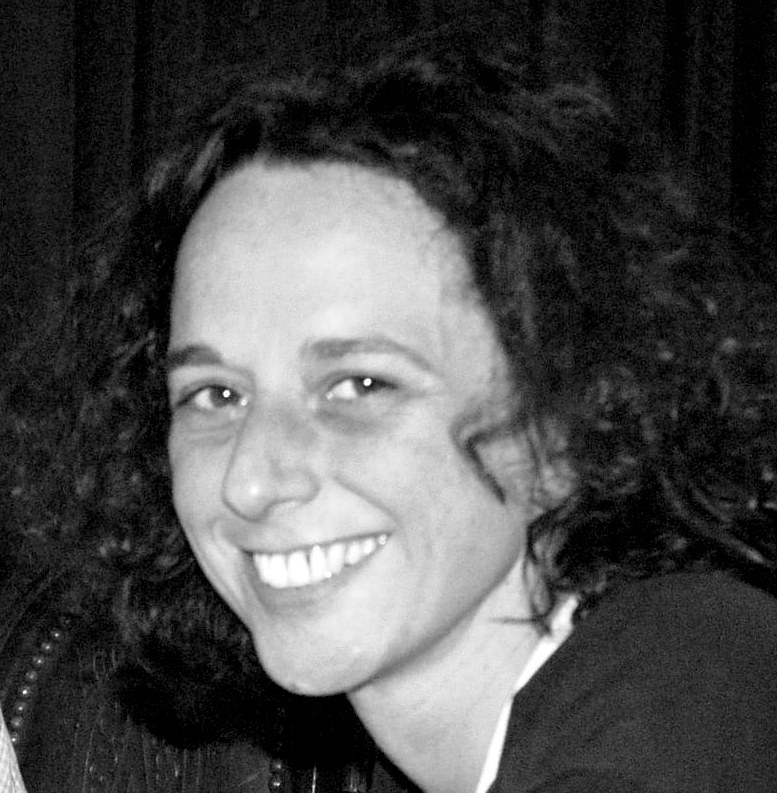 Jocelyne Demengeot and I met at the Institut Pasteur. She returned to France from a postdoctoral period in Boston about
Jocelyne Demengeot and I met at the Institut Pasteur. She returned to France from a postdoctoral period in Boston about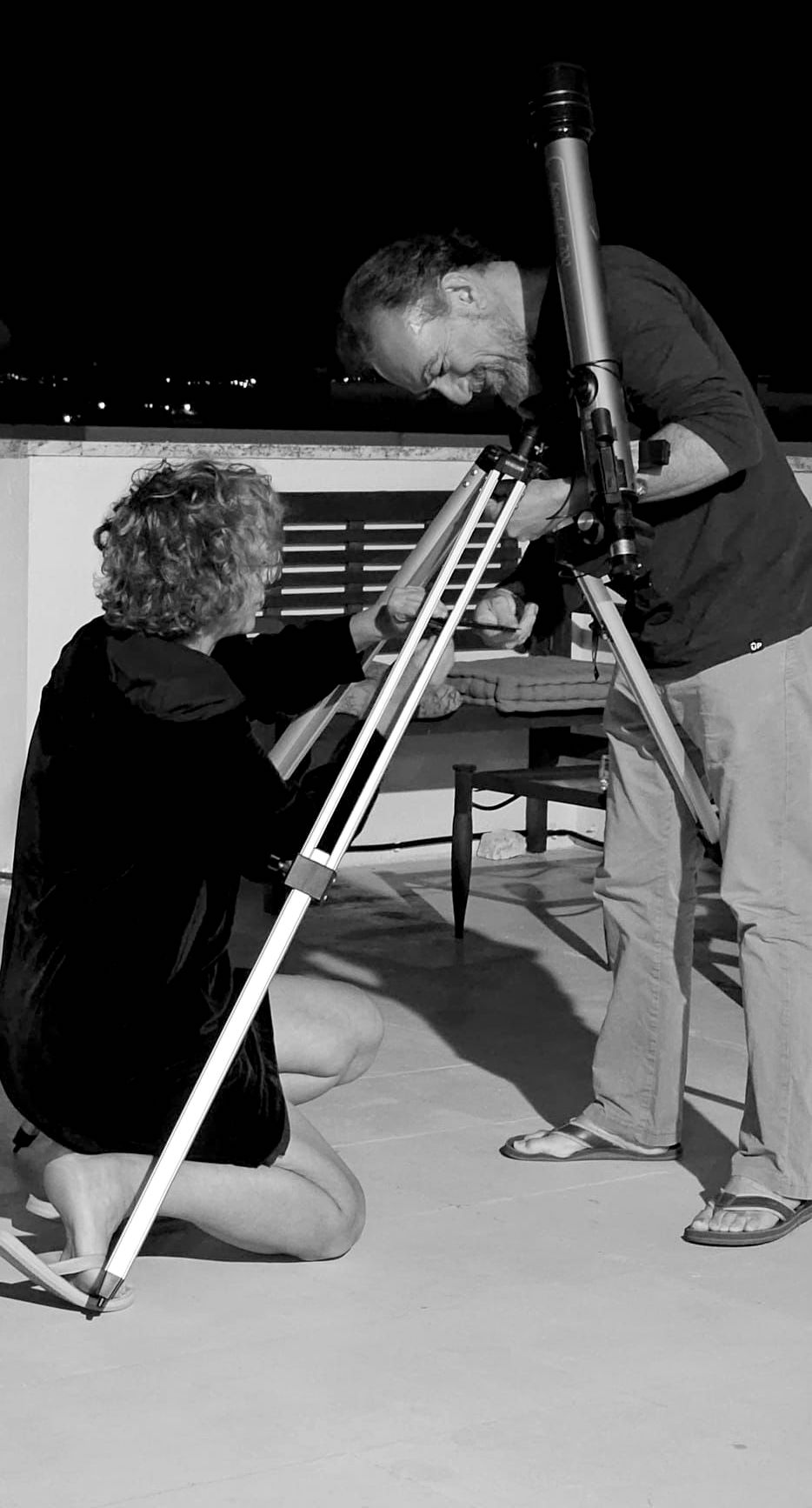  the same time I was wrapping up my PhD thesis. On the hindsight, the fact that we did not get along well immediately is intriguing given that ever since we have enjoyed doing things together—and we really did a lot things together. We were both founding members and actors in the renewal of Instituto Gulbenkian de Ciência around 1998, we shared lab meetings and students, got awarded grants and prizes, coauthored papers, taught courses, served at the direction of the Portuguese Society of Immunology, were consecutively at the helm of the IGC as Deputy director for Science, and we independently decided to step down from the institute, after the FCG announced the merger of IGC with IMM to make what would become the GIMM. Jocelyne is the person I go to to brainstorm about any subject, scientific or personal. It is unclear how much I influenced her, but Jocelyne —the scientist, the colleague and the friend—has shaped who I am. “[We] have seen things you people wouldn’t believe.”
the same time I was wrapping up my PhD thesis. On the hindsight, the fact that we did not get along well immediately is intriguing given that ever since we have enjoyed doing things together—and we really did a lot things together. We were both founding members and actors in the renewal of Instituto Gulbenkian de Ciência around 1998, we shared lab meetings and students, got awarded grants and prizes, coauthored papers, taught courses, served at the direction of the Portuguese Society of Immunology, were consecutively at the helm of the IGC as Deputy director for Science, and we independently decided to step down from the institute, after the FCG announced the merger of IGC with IMM to make what would become the GIMM. Jocelyne is the person I go to to brainstorm about any subject, scientific or personal. It is unclear how much I influenced her, but Jocelyne —the scientist, the colleague and the friend—has shaped who I am. “[We] have seen things you people wouldn’t believe.”
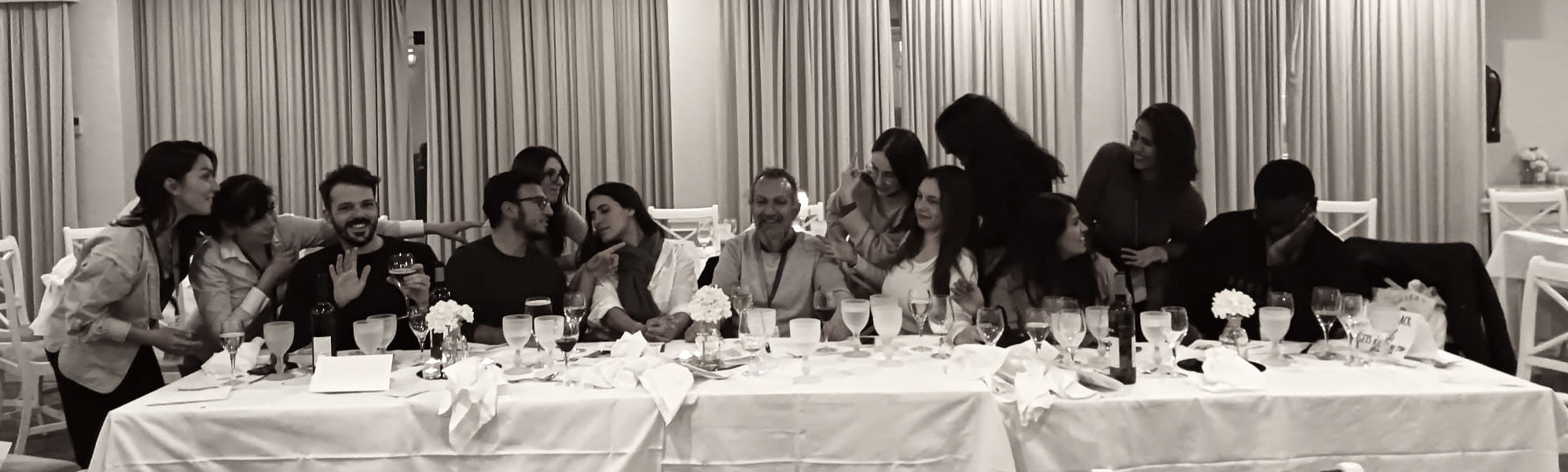 "The Last Supper" enacted by the IBB2021, students admitted to the Integrative Biology and Biomedicine Programme in 2021, during the last student retreat AMeeGuS in which I participated as Programme Director.
"The Last Supper" enacted by the IBB2021, students admitted to the Integrative Biology and Biomedicine Programme in 2021, during the last student retreat AMeeGuS in which I participated as Programme Director.
 Andrés Aldana did his thesis with Alberto Darszon, aiming to develop a logical dynamics model of the acrosome reaction in human sperm. As part of this work, he visited my lab to focus on a specific challenge: using the regulatory network model to explain sperm heterogeneity, particularly, the differences in their ability to undergo spontaneous or induced acrosomal reaction. This required not only linking network attractors to cellular fates, as is common practice, but also defining a biologically meaningful distribution of initial states. Unsatisfied with the standard assumption of equiprobable initial states, we began developing more realistic approaches.
Andrés Aldana did his thesis with Alberto Darszon, aiming to develop a logical dynamics model of the acrosome reaction in human sperm. As part of this work, he visited my lab to focus on a specific challenge: using the regulatory network model to explain sperm heterogeneity, particularly, the differences in their ability to undergo spontaneous or induced acrosomal reaction. This required not only linking network attractors to cellular fates, as is common practice, but also defining a biologically meaningful distribution of initial states. Unsatisfied with the standard assumption of equiprobable initial states, we began developing more realistic approaches.
 Estefanía Muñoz Hoyos completed her studies and PhD at the National University of Colombia, with academic stays at Texas A&M University and the Politecnico di Torino. Her doctoral research focused on the ecohydrology of water- and energy-limited ecosystems. After her PhD, she worked at the World Mosquito Program and then joined the Biology by Numbers Postdoctoral Programme at the Instituto Gulbenkian de Ciência. In the context of this programme, she took part in the classes from the PhD Programme in Integrative Biology. Early on, we discovered a shared curiosity about the role of plant-microbe symbiosis in shaping global ecological dynamics and system robustness. Together, we explored this question using the Daisyworld, a planetary ecology toy model, showing that plant-microbe symbiosis can expand the habitability range of the planet. After IGC, Estefanía joined the Max Planck Institute for Biogeochemistry. She is currently a Marie Curie postdoctoral fellow at CREAF, in Barcelona.
Estefanía Muñoz Hoyos completed her studies and PhD at the National University of Colombia, with academic stays at Texas A&M University and the Politecnico di Torino. Her doctoral research focused on the ecohydrology of water- and energy-limited ecosystems. After her PhD, she worked at the World Mosquito Program and then joined the Biology by Numbers Postdoctoral Programme at the Instituto Gulbenkian de Ciência. In the context of this programme, she took part in the classes from the PhD Programme in Integrative Biology. Early on, we discovered a shared curiosity about the role of plant-microbe symbiosis in shaping global ecological dynamics and system robustness. Together, we explored this question using the Daisyworld, a planetary ecology toy model, showing that plant-microbe symbiosis can expand the habitability range of the planet. After IGC, Estefanía joined the Max Planck Institute for Biogeochemistry. She is currently a Marie Curie postdoctoral fellow at CREAF, in Barcelona.
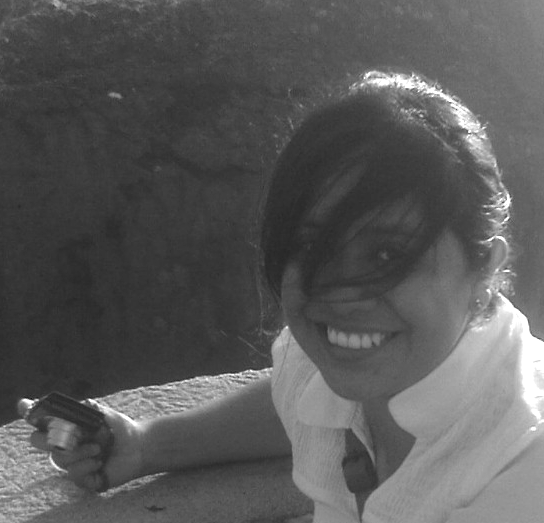 Monserrat Alba Sandoval-Hernández was beginning her PhD with Yvonne Rosenstein when I was on sabbatical at Instituto de Biotecnologia in Cuernavaca. I helped her analyse her data, and we became friends. Later, Monse visited me while working on a draft of her article detailing how CD43 and CD28 co-signalling with the TCR impact T-cell differentiation, depending on the temporal context.
Monserrat Alba Sandoval-Hernández was beginning her PhD with Yvonne Rosenstein when I was on sabbatical at Instituto de Biotecnologia in Cuernavaca. I helped her analyse her data, and we became friends. Later, Monse visited me while working on a draft of her article detailing how CD43 and CD28 co-signalling with the TCR impact T-cell differentiation, depending on the temporal context.
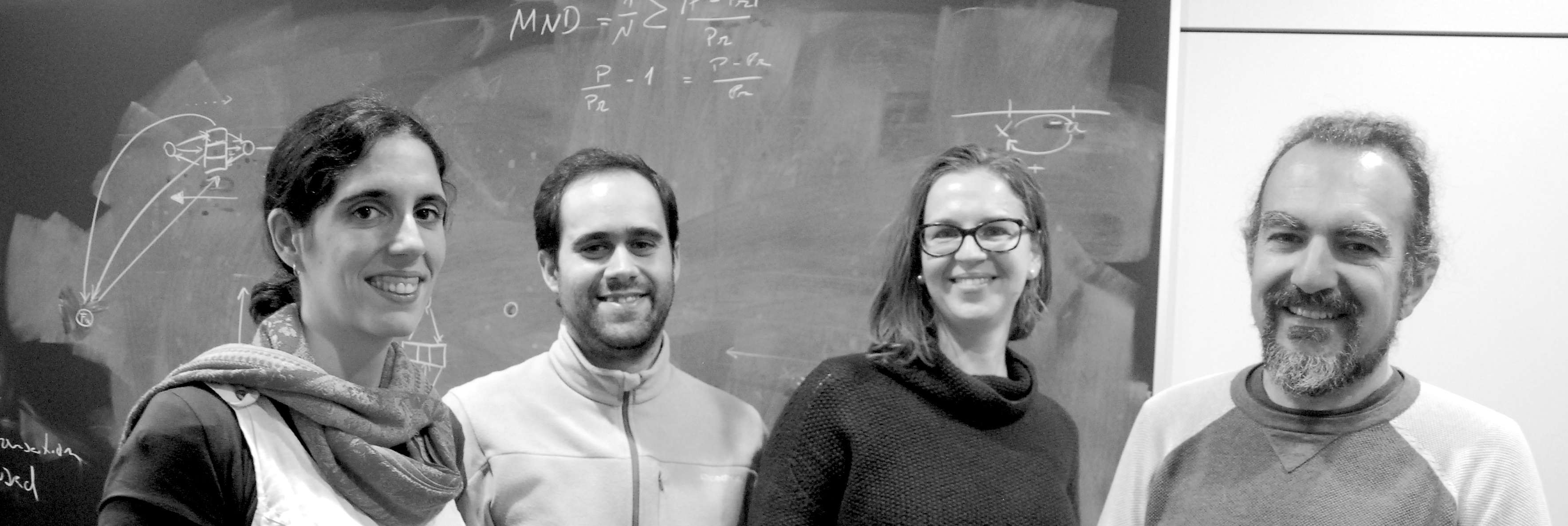 Photo of my research group for the institute's 2018 annual report. From left to right, Delphine, Pedro, Eleonora and me.
Photo of my research group for the institute's 2018 annual report. From left to right, Delphine, Pedro, Eleonora and me.
 Eleonora Tulumello was my last official PhD student. After completing secondary school in the Humanities, she turned to Applied Mathematics, seeking to combine creativity with rigour. During her studies, she developed a strong interest in emergent and collective behaviours and was introduced to research at the Department of Mathematics at the University of Palermo, where she implemented and analysed reaction-diffusion models in ecological contexts. Her work focused on the emergence of self-organised spatio-temporal patterns and the transition to chaotic dynamics. She later joined a short-term research project at the Institute of Biophysics (National Research Council, Palermo), investigating how neural network self-organisation influences odour perception.
Eleonora Tulumello was my last official PhD student. After completing secondary school in the Humanities, she turned to Applied Mathematics, seeking to combine creativity with rigour. During her studies, she developed a strong interest in emergent and collective behaviours and was introduced to research at the Department of Mathematics at the University of Palermo, where she implemented and analysed reaction-diffusion models in ecological contexts. Her work focused on the emergence of self-organised spatio-temporal patterns and the transition to chaotic dynamics. She later joined a short-term research project at the Institute of Biophysics (National Research Council, Palermo), investigating how neural network self-organisation influences odour perception.
Eleonora entered the Integrative Biology and Biomedicine PhD Programme at IGC, choosing to carry out a joint project between my group and Jocelyne Demengeot’s Lymphocyte Physiology Lab. Her goal was to bridge "wet" and "dry" lab approaches, using mathematical models to generate hypotheses for experimental testing. Her thesis explored 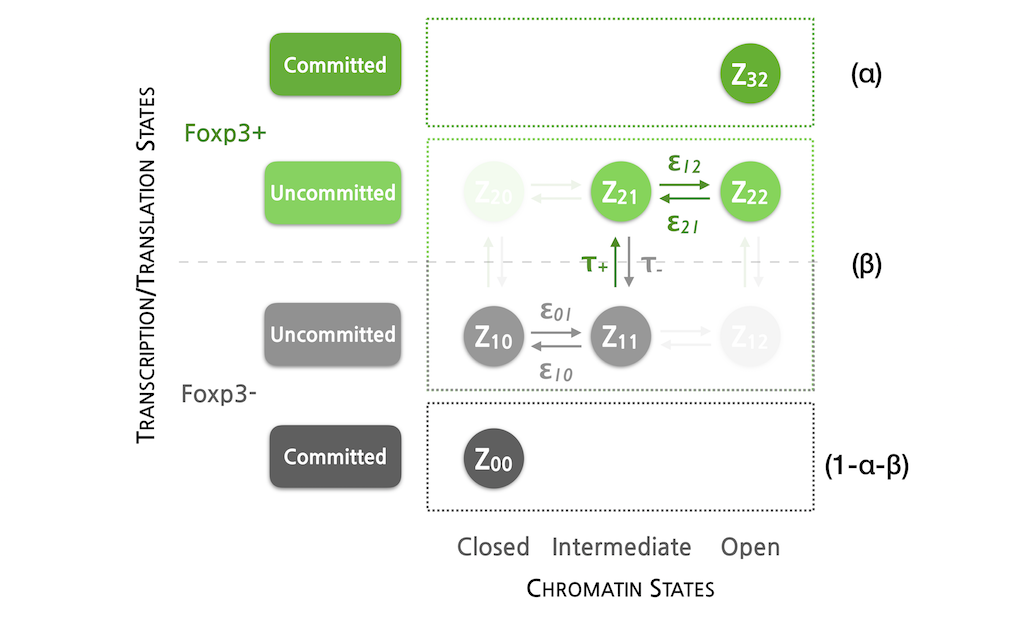 how Foxp3 gene regulation in CD4 T cells, in conjunction with immune population dynamics, contributes to the robust maintenance of natural tolerance. She earned her PhD in July 2020 from NOVA University of Lisbon with the dissertation "Robust tolerance out of volatile regulatory T cells? Lessons from mathematical modelling."
how Foxp3 gene regulation in CD4 T cells, in conjunction with immune population dynamics, contributes to the robust maintenance of natural tolerance. She earned her PhD in July 2020 from NOVA University of Lisbon with the dissertation "Robust tolerance out of volatile regulatory T cells? Lessons from mathematical modelling."
After completing her PhD, Eleonora pursued postgraduate studies in science communication at SISA and is now an inspired and inspiring science communicator, working in the Science Communication and Outreach Office at ITQB NOVA.
 Delphine Pessoa studied Biology for her Bachelor’s and Computational Biology for her Master’s. She joined the IGC to carry out her Master’s research with Gabriela Gomes, working on parameter inference for pneumococcus transmission among day-care attendees and on estimating susceptibility distributions from dose-response experiments. In 2014, she enrolled in the Integrative Biology and Biomedicine PhD Programme.
Delphine Pessoa studied Biology for her Bachelor’s and Computational Biology for her Master’s. She joined the IGC to carry out her Master’s research with Gabriela Gomes, working on parameter inference for pneumococcus transmission among day-care attendees and on estimating susceptibility distributions from dose-response experiments. In 2014, she enrolled in the Integrative Biology and Biomedicine PhD Programme.
 I was struck by her intellect during the statistics course, when she independently derived the Fisher exact test from first principles. I felt privileged when she asked me to supervise her PhD project. Together, we explored a fundamental question: how do biosynthetic processes in cells reliably produce just one product per cell? Delphine received several awards for her poster presentations and earned her PhD in May 2019 from NOVA University of Lisbon with a thesis entitled “How to make a single product per cell? Lessons from X chromosome inactivation and antigen receptor gene recombination.”
I was struck by her intellect during the statistics course, when she independently derived the Fisher exact test from first principles. I felt privileged when she asked me to supervise her PhD project. Together, we explored a fundamental question: how do biosynthetic processes in cells reliably produce just one product per cell? Delphine received several awards for her poster presentations and earned her PhD in May 2019 from NOVA University of Lisbon with a thesis entitled “How to make a single product per cell? Lessons from X chromosome inactivation and antigen receptor gene recombination.”
Delphine is now a Data Scientist at Phrase, after previous roles at Closer Consulting and Unbabel.
 Pedro Silva and I first met during his Master’s in Computational Biology, when he joined my lab for his research project. He produced outstanding work on quantitative imaging of spermatozoa, earning top marks for his dissertation in 2011. Motivated to continue, Pedro secured an individual doctoral fellowship from Fundação para a Ciência e a Tecnologia and began his PhD in collaboration with Alberto Darszon and colleagues at the Instituto de Biotecnología, UNAM, Cuernavaca. Pedro asked how sea urchin spermatozoa navigate in three dimensions to locate conspecific eggs during broadcast spawning. This led him to develop novel image analysis methods for tracking sperm cells in both 2D and 3D.
Pedro Silva and I first met during his Master’s in Computational Biology, when he joined my lab for his research project. He produced outstanding work on quantitative imaging of spermatozoa, earning top marks for his dissertation in 2011. Motivated to continue, Pedro secured an individual doctoral fellowship from Fundação para a Ciência e a Tecnologia and began his PhD in collaboration with Alberto Darszon and colleagues at the Instituto de Biotecnología, UNAM, Cuernavaca. Pedro asked how sea urchin spermatozoa navigate in three dimensions to locate conspecific eggs during broadcast spawning. This led him to develop novel image analysis methods for tracking sperm cells in both 2D and 3D. 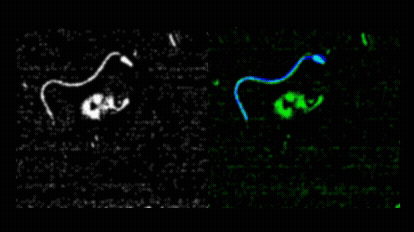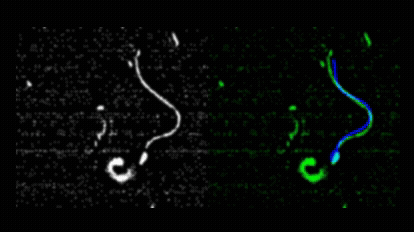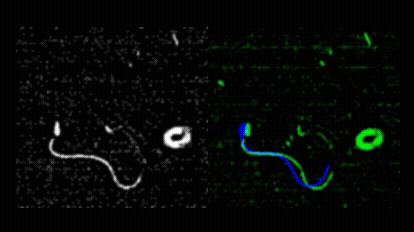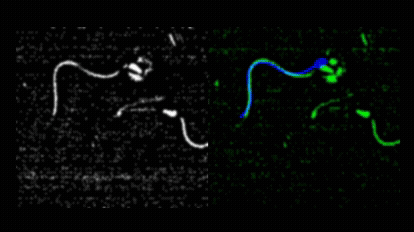 He introduced a mechanical model of spermatozoa to unify image processing, tracking, and trajectory reconstruction into a single nonlinear model-fitting problem.
He introduced a mechanical model of spermatozoa to unify image processing, tracking, and trajectory reconstruction into a single nonlinear model-fitting problem.  He also redesigned the image analysis workflow used in the Corkidi and Darszon labs for 3D microscopy, and performed a meta-analysis of sperm swimming trajectories across species, comparing L. pictus and S. purpuratus under free-swimming and surface-constrained conditions, and integrating data from A. punctulata as well. Pedro earned his PhD in Computational Biology from NOVA University Lisbon in May 2017 with a thesis entitled “Quantitative image analysis of cells using morphodynamical models: Sea urchin spermatozoa as case study.”
He also redesigned the image analysis workflow used in the Corkidi and Darszon labs for 3D microscopy, and performed a meta-analysis of sperm swimming trajectories across species, comparing L. pictus and S. purpuratus under free-swimming and surface-constrained conditions, and integrating data from A. punctulata as well. Pedro earned his PhD in Computational Biology from NOVA University Lisbon in May 2017 with a thesis entitled “Quantitative image analysis of cells using morphodynamical models: Sea urchin spermatozoa as case study.”
He is now a happy Senior Full Stack Team Leader at Talkdesk, after a previous role at Mobiag.
 Marco Louro developed his research project for a Master’s degree in Bioinformatics and Computational Biology in my group, in collaboration with Mónica Dias. With a Bachelor’s degree in Biology, specializing in Evolution and Development, Marco set out to address two interrelated questions: how a cell produces a single daughter centriole from each mother centriole, and how PLK4 concentration determines whether a single or multiple centrioles are formed. He proposed an original and bold hypothesis suggesting that the elongation of the first cartwheel inhibits the formation of additional ones by depleting the shared pool of building blocks. Although initially met with skepticism by both theoreticians and experimentalists, Marco eventually persuaded us with his compelling quantitative arguments and insightful interpretation of the literature. I was deeply impressed by the independent thinking and raw creativity he demonstrated at such an early stage of his career. He received his Master’s degree in Computational Biology from the Faculty of Sciences of the University of Lisbon in November 2016, for his dissertation entitled "A Stochastic Model of Centriole Assembly". He later joined the PhD Programme in Integrative Biology and Biomedicine and carried out his doctoral research under the supervision of Cláudia Bank and Mónica Bettencourt-Dias.
Marco Louro developed his research project for a Master’s degree in Bioinformatics and Computational Biology in my group, in collaboration with Mónica Dias. With a Bachelor’s degree in Biology, specializing in Evolution and Development, Marco set out to address two interrelated questions: how a cell produces a single daughter centriole from each mother centriole, and how PLK4 concentration determines whether a single or multiple centrioles are formed. He proposed an original and bold hypothesis suggesting that the elongation of the first cartwheel inhibits the formation of additional ones by depleting the shared pool of building blocks. Although initially met with skepticism by both theoreticians and experimentalists, Marco eventually persuaded us with his compelling quantitative arguments and insightful interpretation of the literature. I was deeply impressed by the independent thinking and raw creativity he demonstrated at such an early stage of his career. He received his Master’s degree in Computational Biology from the Faculty of Sciences of the University of Lisbon in November 2016, for his dissertation entitled "A Stochastic Model of Centriole Assembly". He later joined the PhD Programme in Integrative Biology and Biomedicine and carried out his doctoral research under the supervision of Cláudia Bank and Mónica Bettencourt-Dias.
 Daniel Priego and I collaborated when he was conducting his PhD research at the Department of Physics at UNAM, Cuernavaca, under the supervision of Gustavo Martinez Mekler and Alberto Darszon. He visited my group at the Instituto Gulbenkian de Ciência several times between 2015 and 2017.The main goal of Daniel’s thesis was to investigate the structure and dynamics of the signalling network that governs the chemotactic response of sea urchin spermatozoa to peptides released by conspecific eggs. This network orchestrates cytosolic Ca²⁺ spike trains, timed to modulate the flagellar bending waves responsible for propulsion and steering. Daniel developed an ordinary differential equation model that thoroughly incorporated existing experimental data, drawing from the literature on each candidate ion channel to identify functional forms and parameter values. This was an Herculean comparative study that grounded experimentally Daniel's modelling efforts. His analysis revealed that two distinct, partially overlapping, combinations of channels —featuring either Ca²⁺ channels CaV or CatSper— could replicate the envelope and temporal dynamics of the observed Ca²⁺ spike trains.
Daniel Priego and I collaborated when he was conducting his PhD research at the Department of Physics at UNAM, Cuernavaca, under the supervision of Gustavo Martinez Mekler and Alberto Darszon. He visited my group at the Instituto Gulbenkian de Ciência several times between 2015 and 2017.The main goal of Daniel’s thesis was to investigate the structure and dynamics of the signalling network that governs the chemotactic response of sea urchin spermatozoa to peptides released by conspecific eggs. This network orchestrates cytosolic Ca²⁺ spike trains, timed to modulate the flagellar bending waves responsible for propulsion and steering. Daniel developed an ordinary differential equation model that thoroughly incorporated existing experimental data, drawing from the literature on each candidate ion channel to identify functional forms and parameter values. This was an Herculean comparative study that grounded experimentally Daniel's modelling efforts. His analysis revealed that two distinct, partially overlapping, combinations of channels —featuring either Ca²⁺ channels CaV or CatSper— could replicate the envelope and temporal dynamics of the observed Ca²⁺ spike trains. 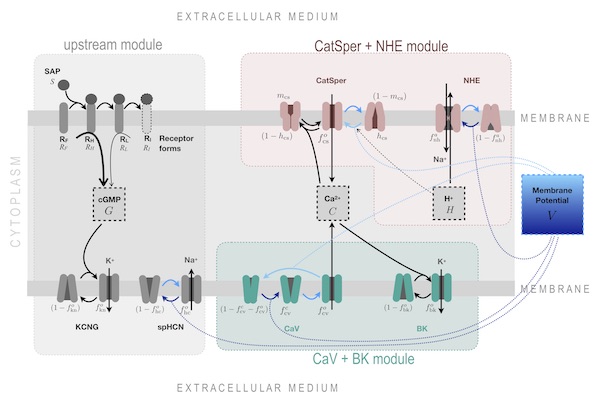 Seeking to understand how different network architectures could yield similar outputs, Daniel carried out a detailed bifurcation and modular analysis of the model variants. This approach provided the insight he needed and helped identify key distinguishing features that could be tested experimentally. Based on these predictions, he concluded in favour of the model featuring the CatSper channel.
Seeking to understand how different network architectures could yield similar outputs, Daniel carried out a detailed bifurcation and modular analysis of the model variants. This approach provided the insight he needed and helped identify key distinguishing features that could be tested experimentally. Based on these predictions, he concluded in favour of the model featuring the CatSper channel.
Daniel received his doctoral degree from UNAM in 2018. He is currently a postdoctoral researcher in the Department of Biology at the University of Kentucky, Lexington, USA.
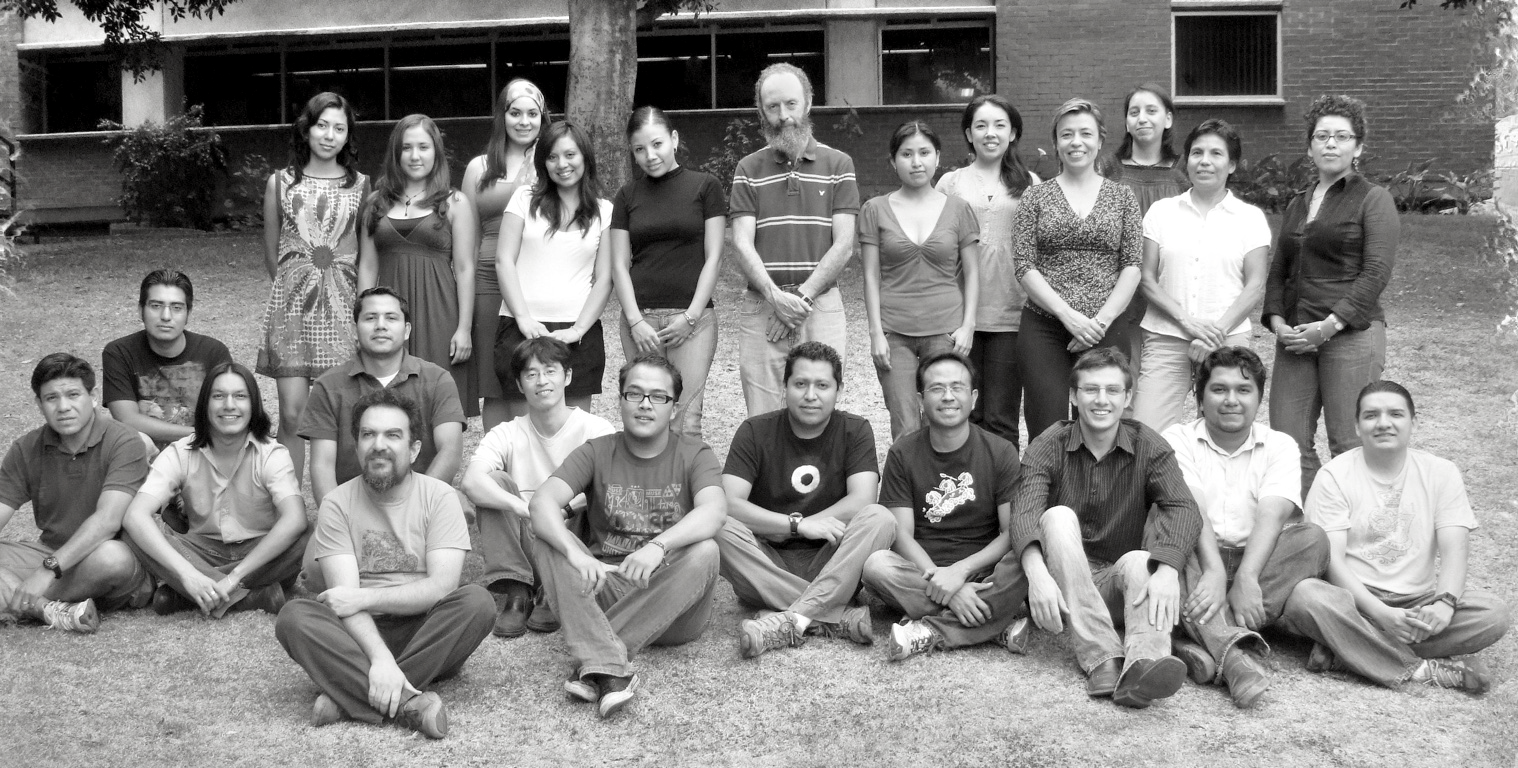 Group photo of Alberto Darszon's laboratory at the Instituto de Biotecnologia in Cuernavaca, taken during my sabbatical in 2010.
Group photo of Alberto Darszon's laboratory at the Instituto de Biotecnologia in Cuernavaca, taken during my sabbatical in 2010.
 Arturo Pimentel and I met when he was a PhD student working with Gabriel Corkidi at the Instituto de Biotecnología in Cuernavaca. Arturo and Gabriel were developing an ingenious microscopy system for Alberto Darszon’s lab, aimed at capturing the 3D trajectories of fast-swimming spermatozoa. Their setup combined a piezoelectric device to move the objective’s focal plane rapidly through a volume with an ultra-fast camera acquiring thousands of frames per second. At that early stage, they faced significant challenges, the most proeminent being the absence of information on the z-coordinate of each frame. Joining their effort, I proposed to use pairwise correlations to aligm frames by z-coordinate, and worked with Arturo to implement this solution as an algorithm. Arturo visited me at the Instituto Gulbenkian de Ciência, where we fine-tuned the details of the method.
After concluding his thesis, Arturo joined the National Microscopy Laboratory as a technician. Arturo and I have recently reconnected, through our shared commitment to the quantitative education of young researchers. Arturo is my local collaborator in the Systems Biology course I teach to students of several of UNAM’s graduate programmes. He’s my go-to person for anything microscopy instrumentation-related and an unrelenting guide when it comes to navigating UNAM’s administration. He is also diligently completing my education in Mexican real politics and cuisine. Arturo’s stubbornness is only surpassed by my own, and I’ve come to admire his instinct for walking that fine line between wisdom and cynicism, never slipping into paralysis.
Arturo Pimentel and I met when he was a PhD student working with Gabriel Corkidi at the Instituto de Biotecnología in Cuernavaca. Arturo and Gabriel were developing an ingenious microscopy system for Alberto Darszon’s lab, aimed at capturing the 3D trajectories of fast-swimming spermatozoa. Their setup combined a piezoelectric device to move the objective’s focal plane rapidly through a volume with an ultra-fast camera acquiring thousands of frames per second. At that early stage, they faced significant challenges, the most proeminent being the absence of information on the z-coordinate of each frame. Joining their effort, I proposed to use pairwise correlations to aligm frames by z-coordinate, and worked with Arturo to implement this solution as an algorithm. Arturo visited me at the Instituto Gulbenkian de Ciência, where we fine-tuned the details of the method.
After concluding his thesis, Arturo joined the National Microscopy Laboratory as a technician. Arturo and I have recently reconnected, through our shared commitment to the quantitative education of young researchers. Arturo is my local collaborator in the Systems Biology course I teach to students of several of UNAM’s graduate programmes. He’s my go-to person for anything microscopy instrumentation-related and an unrelenting guide when it comes to navigating UNAM’s administration. He is also diligently completing my education in Mexican real politics and cuisine. Arturo’s stubbornness is only surpassed by my own, and I’ve come to admire his instinct for walking that fine line between wisdom and cynicism, never slipping into paralysis.
 Adan Guerrero and I started working on the quantification of sea urchin sperm chemotaxis back in 2008. We had met the year before, during a three-weeks workshop organised by Denis Thieffry and Gustavo Mekler at the Centro Internacional de Ciências in Cuernavaca. I had been trying to use a lymphocyte morphodynamics model as a tool to analyse imaging data and estimating model parameters, but the degrees of freedom of lymphocyte shape was hard to handle. The sea urchin spermatozoa was the ideal system to test such morphodynamical model-based image analysis as these cells swim confined to planar water interfaces. When I met Adan and Alberto Darszon, who were grappling with quantitative sperm imaging, I immediately saw an opportunity for collaboration. The seminal model, I developed with Adan would later be improved and extended to three dimensions by Pedro Silva. Meanwhile, the collaboration with Adan and Alberto broadned to other research(ers). Returning to Adan, he stood out from the start as a highly impressive PhD student, and over time, it became clear just how extraordinary he was. Driven by intellectual curiosity and strong focus, Adan acquired a broad and solid set of skills with the goal better understanding and exploring his data. He became the go-to person in his lab for tweaking and custom-building microscopy setups, eventually playing some role in developing instrumentation for 3D tracking of sperm trajectories. He initiated collaborations with physicists to deepen the theoretical understanding of sperm signalling networks using boolean networks. With me, he wanted to improve the quantification and automation of image analysis. Recognising his own limitations in statistics, imaging, and programming, Adan took a university course in computer vision after hours, voluntarily and without letting it interfere with his lab work. That investment paid off. By the time he completed his thesis, Adan had become impressively self-sufficient: fluent in multiple programming languages, capable of writing his own image analysis software, and able to channel his time and energy into more insightful and productive analyses.
Adan was the perfect team mate during my sabbatical in Cuernavaca in 2010. We spent most our time working or discussing science, and outside the lab, he introduced me to Mexican culture, food, and way of life, driving me around in his miraculously functioning Vochito. After a postdoc with Mónica Dias at the Instituto Gulbenkian de Ciência, Adan returned to Mexico and is now a researcher at the National Microscopy Laboratory, having been awarded recently the Prémio National de Investigação Científica. In a fitting turn of events, he is today the director of the Centro Internacional de Ciencias, where we first met.
Adan Guerrero and I started working on the quantification of sea urchin sperm chemotaxis back in 2008. We had met the year before, during a three-weeks workshop organised by Denis Thieffry and Gustavo Mekler at the Centro Internacional de Ciências in Cuernavaca. I had been trying to use a lymphocyte morphodynamics model as a tool to analyse imaging data and estimating model parameters, but the degrees of freedom of lymphocyte shape was hard to handle. The sea urchin spermatozoa was the ideal system to test such morphodynamical model-based image analysis as these cells swim confined to planar water interfaces. When I met Adan and Alberto Darszon, who were grappling with quantitative sperm imaging, I immediately saw an opportunity for collaboration. The seminal model, I developed with Adan would later be improved and extended to three dimensions by Pedro Silva. Meanwhile, the collaboration with Adan and Alberto broadned to other research(ers). Returning to Adan, he stood out from the start as a highly impressive PhD student, and over time, it became clear just how extraordinary he was. Driven by intellectual curiosity and strong focus, Adan acquired a broad and solid set of skills with the goal better understanding and exploring his data. He became the go-to person in his lab for tweaking and custom-building microscopy setups, eventually playing some role in developing instrumentation for 3D tracking of sperm trajectories. He initiated collaborations with physicists to deepen the theoretical understanding of sperm signalling networks using boolean networks. With me, he wanted to improve the quantification and automation of image analysis. Recognising his own limitations in statistics, imaging, and programming, Adan took a university course in computer vision after hours, voluntarily and without letting it interfere with his lab work. That investment paid off. By the time he completed his thesis, Adan had become impressively self-sufficient: fluent in multiple programming languages, capable of writing his own image analysis software, and able to channel his time and energy into more insightful and productive analyses.
Adan was the perfect team mate during my sabbatical in Cuernavaca in 2010. We spent most our time working or discussing science, and outside the lab, he introduced me to Mexican culture, food, and way of life, driving me around in his miraculously functioning Vochito. After a postdoc with Mónica Dias at the Instituto Gulbenkian de Ciência, Adan returned to Mexico and is now a researcher at the National Microscopy Laboratory, having been awarded recently the Prémio National de Investigação Científica. In a fitting turn of events, he is today the director of the Centro Internacional de Ciencias, where we first met.
 My research group in 2011. From left to right, Thiago, Pedro, Macedo, me, Danesh and Tom.
My research group in 2011. From left to right, Thiago, Pedro, Macedo, me, Danesh and Tom.
 My collaboration and friendship with the late Claudine Chaouiya dated back to the time when we worked together with Aurelien Naldi and Denis Thieffry on the logical modelling of the gene regulatory networks that control the differentiation of CD4+ T cells. She later joined the Instituto Gulbenkian de Ciência, to help me with the PhD Programme in Computational Biology and to become an independent research with her own group. We collaborated in different aspects of network biology, namely in the development of new formal tools to deal with the challenges that multicellular systems pose to gene regulatory network theory. She eventually returned to Marseille to the Mathematics Institute of Marseille. Claudine left us saddly in 2025. Everytime I get served an expresso with a sugar bag on the side, I warmly remember Claudine smiling and saying "Para doce basta a vida!"
My collaboration and friendship with the late Claudine Chaouiya dated back to the time when we worked together with Aurelien Naldi and Denis Thieffry on the logical modelling of the gene regulatory networks that control the differentiation of CD4+ T cells. She later joined the Instituto Gulbenkian de Ciência, to help me with the PhD Programme in Computational Biology and to become an independent research with her own group. We collaborated in different aspects of network biology, namely in the development of new formal tools to deal with the challenges that multicellular systems pose to gene regulatory network theory. She eventually returned to Marseille to the Mathematics Institute of Marseille. Claudine left us saddly in 2025. Everytime I get served an expresso with a sugar bag on the side, I warmly remember Claudine smiling and saying "Para doce basta a vida!"
 Danesh Tarapore joined my lab as a postdoctoral researcher in 2010 to work in a collaborative project with Pedro Lima from Instituto Superior Técnico and Anders Lynn Christensen from ISCTE in Lisbon. We were fortunate to have coopted Danesh to our project. He was trained in computer science and had a strong interest in interdisciplinary research. His work spaned computer science, robotics, and biology. He earned his PhD in Life Sciences from the University of Lausanne, Switzerland, where he investigated how genetic and social factors influence division of labour in social insect colonies, and also examined the performance impact of different task allocation mechanisms within colonies.
Our project, BioInstBots aimed to create a theory of collective robotic systems inspired both in biology and institutional economics. Danesh successfully explored how to equip these systems with distributed immune mechanisms for detecting and responding to unexpected faults and anomalies. He also developed an efficient agent-based simulation framework capable of modelling both cellular gene regulatory networks dynamics within the cell and the supra-cellular dynamics.
Danesh Tarapore joined my lab as a postdoctoral researcher in 2010 to work in a collaborative project with Pedro Lima from Instituto Superior Técnico and Anders Lynn Christensen from ISCTE in Lisbon. We were fortunate to have coopted Danesh to our project. He was trained in computer science and had a strong interest in interdisciplinary research. His work spaned computer science, robotics, and biology. He earned his PhD in Life Sciences from the University of Lausanne, Switzerland, where he investigated how genetic and social factors influence division of labour in social insect colonies, and also examined the performance impact of different task allocation mechanisms within colonies.
Our project, BioInstBots aimed to create a theory of collective robotic systems inspired both in biology and institutional economics. Danesh successfully explored how to equip these systems with distributed immune mechanisms for detecting and responding to unexpected faults and anomalies. He also developed an efficient agent-based simulation framework capable of modelling both cellular gene regulatory networks dynamics within the cell and the supra-cellular dynamics.
He is currently an Assistant Professor in the Agents, Interaction and Complexity Group, Electronics and Computer Science, at the University of Southampton. Visit his personal webpage here.
 Tom Weber worked with Michal Or-Guil and me, bringing to the collaboration the full strength of his formal training in physics. He was a highly independent PhD student who not only defined his own research objectives but also chose the approaches to pursue them. Tom welcomed criticism, but he was never one to accept a suggestion not to follow a particular route without testing it for himself. This attitude made him explore a very broad range of avenues, some shorter than others, but that enriched his knowledge and skill sets. Motivated by questions around proliferation and selection in germinal centres, he turned his focus to quantifying cell cycle parameters and produced a very original and rigorous body of work. He proposed a new stochastic model for the temporal organisation of the cell cycle that accounted for both mechanistic and stochastic features of the G1, S, G2, and mitosis phases, showing that these could be quantified using flow cytometry data. Tom made an outstanding contribution to the challenge of model identifiability and parameter estimation by designing a new experimental protocol based on dual pulses of nucleoside analogues. This protocol emerged directly from his theoretical work in a fruitful moment of interplay between modelling and experimentation. After his PhD, Tom moved on to work as a postdoc with Ken Duffy in Dublin and after that with Phil Hodgkin in Melbourne.
Tom Weber worked with Michal Or-Guil and me, bringing to the collaboration the full strength of his formal training in physics. He was a highly independent PhD student who not only defined his own research objectives but also chose the approaches to pursue them. Tom welcomed criticism, but he was never one to accept a suggestion not to follow a particular route without testing it for himself. This attitude made him explore a very broad range of avenues, some shorter than others, but that enriched his knowledge and skill sets. Motivated by questions around proliferation and selection in germinal centres, he turned his focus to quantifying cell cycle parameters and produced a very original and rigorous body of work. He proposed a new stochastic model for the temporal organisation of the cell cycle that accounted for both mechanistic and stochastic features of the G1, S, G2, and mitosis phases, showing that these could be quantified using flow cytometry data. Tom made an outstanding contribution to the challenge of model identifiability and parameter estimation by designing a new experimental protocol based on dual pulses of nucleoside analogues. This protocol emerged directly from his theoretical work in a fruitful moment of interplay between modelling and experimentation. After his PhD, Tom moved on to work as a postdoc with Ken Duffy in Dublin and after that with Phil Hodgkin in Melbourne.
 Tiago Macedo was admitted to the PhD Programme in Computational Biology. The programme funders had imposed a peculiar rule: non-national students like Tiago were required to carry out their research in a Portuguese laboratory, whereas Portuguese nationals could go anywhere in the world for their thesis work. Tiago chose to work with me on modelling zebrafish epiboly, initially in collaboration with a colleague who unfortunately left the institute early in the project. Despite this setback, Tiago produced beautiful work. He built on a framework to model cell morphodynamics in a grid-free manner in which the cell boundary was treated as edge connected vertices redesigning it in a computationally efficient and physically sound manner. He used a pattern-forming reaction-diffusion system to control protrusion and retraction activities, generating cell deformation and motility patterns that reproduced those observed in lymphocytes. I was always mesmerised by Tiago’s simulated cells that looked self-driven and very much “alive.” He also applied the model to revisit Albrecht-Buehler's classic analysis of rotational or mirror symmetric trajectories of sister cells, confirming that what had long been seen as meaningful patterns could, in fact, arise from randomness. Tiago, however, struggled with writing about his work. Although the thesis was never brought to completion, he left a lasting impression. He eventually joined the bioinformatics service unit for a few years before moving to the private sector. A passionate maker, he applied his skills in 3D printing to build working microscopes adapted for mobile phones—a highlight for visitors during open days— and devised automated pipetting systems with the same inventive spirit.
Tiago Macedo was admitted to the PhD Programme in Computational Biology. The programme funders had imposed a peculiar rule: non-national students like Tiago were required to carry out their research in a Portuguese laboratory, whereas Portuguese nationals could go anywhere in the world for their thesis work. Tiago chose to work with me on modelling zebrafish epiboly, initially in collaboration with a colleague who unfortunately left the institute early in the project. Despite this setback, Tiago produced beautiful work. He built on a framework to model cell morphodynamics in a grid-free manner in which the cell boundary was treated as edge connected vertices redesigning it in a computationally efficient and physically sound manner. He used a pattern-forming reaction-diffusion system to control protrusion and retraction activities, generating cell deformation and motility patterns that reproduced those observed in lymphocytes. I was always mesmerised by Tiago’s simulated cells that looked self-driven and very much “alive.” He also applied the model to revisit Albrecht-Buehler's classic analysis of rotational or mirror symmetric trajectories of sister cells, confirming that what had long been seen as meaningful patterns could, in fact, arise from randomness. Tiago, however, struggled with writing about his work. Although the thesis was never brought to completion, he left a lasting impression. He eventually joined the bioinformatics service unit for a few years before moving to the private sector. A passionate maker, he applied his skills in 3D printing to build working microscopes adapted for mobile phones—a highlight for visitors during open days— and devised automated pipetting systems with the same inventive spirit.
 Thiago Guzella began working on artificial immune systems while still an undergraduate in Electrical and Electronic Engineering in the Universidade Federal de Minas Gerais in Brazil. We first met at an Artificial Immune Systems meeting in Santos, Brazil, where he impressively presented no fewer than three independent papers. Encouraged by Tomas Mota Santos, former Dean of UFMG and a longtime friend of the Instituto Gulbenkian de Ciência, who recognised his potential, Thiago joined the institute’s PhD Programme and, after completing the course work, chose to work with me on modelling T helper cell differentiation. However, after interacting with other students in my lab, he realised that the questions that drove him were experimental in nature and not easily addressed through theoretical modelling. When he shared this insight, I suggested he find a co-PI to guide the experimental side of his work and Vasco Barreto joined us. The result was a truly interdisciplinary thesis. Thiago spent full days at the bench and full nights with theory (yes, he didn’t sleep). He was interested in the nature of cell-to-cell heterogeneity, particularly in protein copy number. Dissatisfied with superficial answers (semiquantitative at best), he set out to quantify this variability. Experimentally, he sorted naive cells from the high and low tails of TCR expression profiles and tracked their dynamics. Theoretically, he developed a stochastic framework that allowed him to distinguish how much of the observed variance was stable versus volatile, and to estimate the relaxation time of the latter unstable component. He applied this framework to both polyclonal populations with genetically diverse TCRs and transgenic populations with identical TCRs, demonstrating that even in the latter, there was a detectable, non-negligible variance in TCR expression that called for explanation. Thiago defended his thesis in 2017 and then moved on to work in quantitative genetics with Henrique Teotónio at the École Normale Supérieure in Paris. Like many computational biologists, Thiago ultimately found that industry offered more stimulating and rewarding challenges than academia. He is now a Senior Data Scientist at ASML in Amsterdam.
Thiago Guzella began working on artificial immune systems while still an undergraduate in Electrical and Electronic Engineering in the Universidade Federal de Minas Gerais in Brazil. We first met at an Artificial Immune Systems meeting in Santos, Brazil, where he impressively presented no fewer than three independent papers. Encouraged by Tomas Mota Santos, former Dean of UFMG and a longtime friend of the Instituto Gulbenkian de Ciência, who recognised his potential, Thiago joined the institute’s PhD Programme and, after completing the course work, chose to work with me on modelling T helper cell differentiation. However, after interacting with other students in my lab, he realised that the questions that drove him were experimental in nature and not easily addressed through theoretical modelling. When he shared this insight, I suggested he find a co-PI to guide the experimental side of his work and Vasco Barreto joined us. The result was a truly interdisciplinary thesis. Thiago spent full days at the bench and full nights with theory (yes, he didn’t sleep). He was interested in the nature of cell-to-cell heterogeneity, particularly in protein copy number. Dissatisfied with superficial answers (semiquantitative at best), he set out to quantify this variability. Experimentally, he sorted naive cells from the high and low tails of TCR expression profiles and tracked their dynamics. Theoretically, he developed a stochastic framework that allowed him to distinguish how much of the observed variance was stable versus volatile, and to estimate the relaxation time of the latter unstable component. He applied this framework to both polyclonal populations with genetically diverse TCRs and transgenic populations with identical TCRs, demonstrating that even in the latter, there was a detectable, non-negligible variance in TCR expression that called for explanation. Thiago defended his thesis in 2017 and then moved on to work in quantitative genetics with Henrique Teotónio at the École Normale Supérieure in Paris. Like many computational biologists, Thiago ultimately found that industry offered more stimulating and rewarding challenges than academia. He is now a Senior Data Scientist at ASML in Amsterdam.
 José Afonso Guerra joined my lab as a volunteer apprentice while I was developing a lattice-free computational framework to model mammalian cell morphodynamics and he embraced this effort. The initial idea for his internship was for him to run simulations, helping to test the software while learning about the underlying algorithms and how they were implemented. But, unexpectedly for someone in a voluntary role, Afonso quickly began to formulate his own ideas and develop a sharp critical perspective on the work. His performance was so impressive that I soon entrusted him with developing and optimising algorithms and writing new simulation modules. He later earned a place in the PhD Programme in Computational Biology and went on to do his doctoral research on the genomics and evolution of microRNAs with Anton Enright at EMBL-EBI in Cambridge.
José Afonso Guerra joined my lab as a volunteer apprentice while I was developing a lattice-free computational framework to model mammalian cell morphodynamics and he embraced this effort. The initial idea for his internship was for him to run simulations, helping to test the software while learning about the underlying algorithms and how they were implemented. But, unexpectedly for someone in a voluntary role, Afonso quickly began to formulate his own ideas and develop a sharp critical perspective on the work. His performance was so impressive that I soon entrusted him with developing and optimising algorithms and writing new simulation modules. He later earned a place in the PhD Programme in Computational Biology and went on to do his doctoral research on the genomics and evolution of microRNAs with Anton Enright at EMBL-EBI in Cambridge.
 Íris Vilares joined my group at the Instituto Gulbenkian de Ciência as Master student, in 2007. After studying Biology at the Faculty of Sciences of the University of Lisbon, she went on to a Master in Mathematics Applied to the Biological Sciences at the Instituto Superior de Agronomia in Lisbon. She developed her research project towards a Master dissertation with me, being awarded one of the very rare Master fellowships of the Fundação para a Ciência e a Tecnologia from the Portuguese Government. Her research project aimed to better understand the interplay between the immune system and comensal bacteria, building on some ideas previously developed at the group on the role of subclinical infections in the establishment of self-tolerance mediated by regulatory T cells.
Íris Vilares joined my group at the Instituto Gulbenkian de Ciência as Master student, in 2007. After studying Biology at the Faculty of Sciences of the University of Lisbon, she went on to a Master in Mathematics Applied to the Biological Sciences at the Instituto Superior de Agronomia in Lisbon. She developed her research project towards a Master dissertation with me, being awarded one of the very rare Master fellowships of the Fundação para a Ciência e a Tecnologia from the Portuguese Government. Her research project aimed to better understand the interplay between the immune system and comensal bacteria, building on some ideas previously developed at the group on the role of subclinical infections in the establishment of self-tolerance mediated by regulatory T cells.
 Photo taken at the top of the Serra da Estrela during the lab retreat in 2006. From left to right, Luciana, Rui, Daniel, Nuno (on the floor), Tiago, me, and Carline.
Photo taken at the top of the Serra da Estrela during the lab retreat in 2006. From left to right, Luciana, Rui, Daniel, Nuno (on the floor), Tiago, me, and Carline.
 Luciana Itida Ferrari joined my group from 2006 to 2007 through a doutorado sanduíche programme, aiming to deepen her training in the computational analysis of biological data. During her time with us, she developed, with the help of Rui Gardner, a computational framework to analyse data generated by a new technique for estimating T cell diversity in human samples, which was then being developed be colleagues at the Mayo Clinic. She received her PhD in Systems Engineering and Computation from the Universidade Federal do Rio de Janeiro in 2008 and is currently an Associate Professor at the Universidade Federal do Espírito Santo.
Luciana Itida Ferrari joined my group from 2006 to 2007 through a doutorado sanduíche programme, aiming to deepen her training in the computational analysis of biological data. During her time with us, she developed, with the help of Rui Gardner, a computational framework to analyse data generated by a new technique for estimating T cell diversity in human samples, which was then being developed be colleagues at the Mayo Clinic. She received her PhD in Systems Engineering and Computation from the Universidade Federal do Rio de Janeiro in 2008 and is currently an Associate Professor at the Universidade Federal do Espírito Santo.
 Daniel Ferreira joined my lab for an internship in 2005. A psychologist by training, he had developed an interest in computational neuroscience. We worked together on a simple model of C. elegans morphodynamics which was more a vehicle for Daniel to learn coding and simulation than to tackle an original scientific question. He was later accepted into the second edition of the PhD Programme in Computational Biology and went on to pursue his doctoral research in New York. Sadly, for all of us who cherished his camaraderie, his unexpected, unguarded, and slightly unsettling smile, and his maddeningly solid arguments, Daniel left us far too soon. A few posthumous papers remain as testament to his brief scientific journey.
Daniel Ferreira joined my lab for an internship in 2005. A psychologist by training, he had developed an interest in computational neuroscience. We worked together on a simple model of C. elegans morphodynamics which was more a vehicle for Daniel to learn coding and simulation than to tackle an original scientific question. He was later accepted into the second edition of the PhD Programme in Computational Biology and went on to pursue his doctoral research in New York. Sadly, for all of us who cherished his camaraderie, his unexpected, unguarded, and slightly unsettling smile, and his maddeningly solid arguments, Daniel left us far too soon. A few posthumous papers remain as testament to his brief scientific journey.
 Carline van den Dool joined my group for a international internship after completing her degree in Biology at the University of Utrecht in 2005. She developed an individual-oriented simulation of the motility and interactions among regulatory T cells,
conventional T cells, and APCs in vitro. This simulation was able to reproduce qualitatively the cell motility and cluster formation as imaged
in cell cultures by confocal microscopy.
Carline van den Dool joined my group for a international internship after completing her degree in Biology at the University of Utrecht in 2005. She developed an individual-oriented simulation of the motility and interactions among regulatory T cells,
conventional T cells, and APCs in vitro. This simulation was able to reproduce qualitatively the cell motility and cluster formation as imaged
in cell cultures by confocal microscopy. 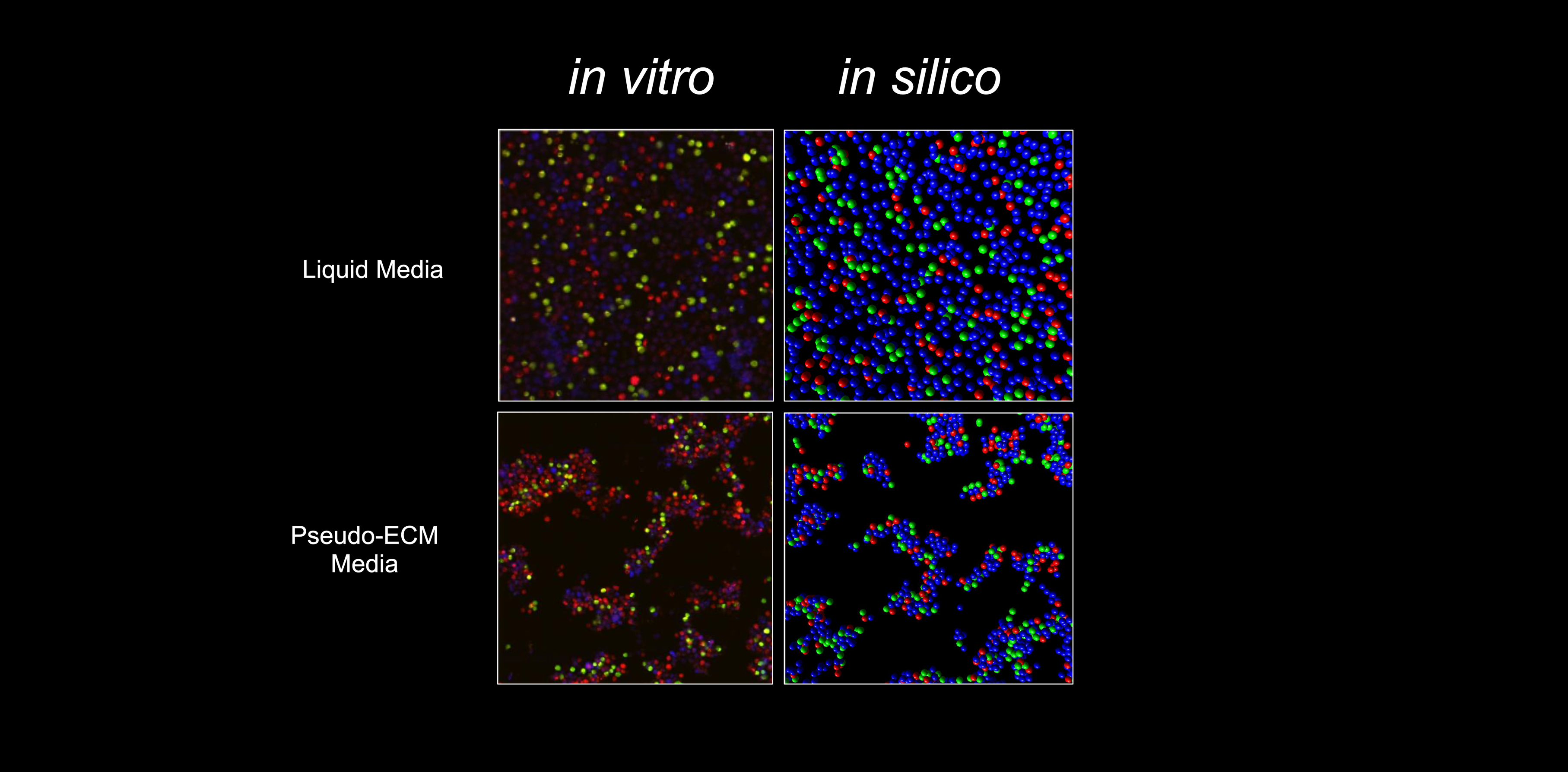 Carline returned to the Netherlands to conduct her Ph.D. research studying the spread of influenza epidemics in hospitals and homes for the elderly in a project that involved both modelling and field work. She left the academia and is currently an event manager and travel organiser
Carline returned to the Netherlands to conduct her Ph.D. research studying the spread of influenza epidemics in hospitals and homes for the elderly in a project that involved both modelling and field work. She left the academia and is currently an event manager and travel organiser
 The group went to the European Conference in Mathematical and Theoretical Biology held in Dresden in 2005. In the photo, Nuno, Ramiro, Thiago and Rui are trying to display a lognormal distribution.
The group went to the European Conference in Mathematical and Theoretical Biology held in Dresden in 2005. In the photo, Nuno, Ramiro, Thiago and Rui are trying to display a lognormal distribution.
 Ramiro Magno joined my group when he was concluding his studies in Biology at the Faculty of Sciences of the University of Lisbon
to prepare his final project on modelling pollen tube growth and navigation with Jose Feijo and me, back in 2004. We asked the question: How are the processes of cell morphogenesis, growth and chemotropism coordinated in a pollen tube? Ramiro developed a simple and computationally efficient algorithm to generate
the growing shape of a pollen tube, based on simple geometrical principles and on the co-regulation of cell-volume expansion and cell-wall extension.
This model was able to reproduce the polarized growth of the pollen tube,
and the normal chemotropism these cells show towards chemoattractants, as well as some abnormal behaviours.
Ramiro Magno joined my group when he was concluding his studies in Biology at the Faculty of Sciences of the University of Lisbon
to prepare his final project on modelling pollen tube growth and navigation with Jose Feijo and me, back in 2004. We asked the question: How are the processes of cell morphogenesis, growth and chemotropism coordinated in a pollen tube? Ramiro developed a simple and computationally efficient algorithm to generate
the growing shape of a pollen tube, based on simple geometrical principles and on the co-regulation of cell-volume expansion and cell-wall extension.
This model was able to reproduce the polarized growth of the pollen tube,
and the normal chemotropism these cells show towards chemoattractants, as well as some abnormal behaviours.
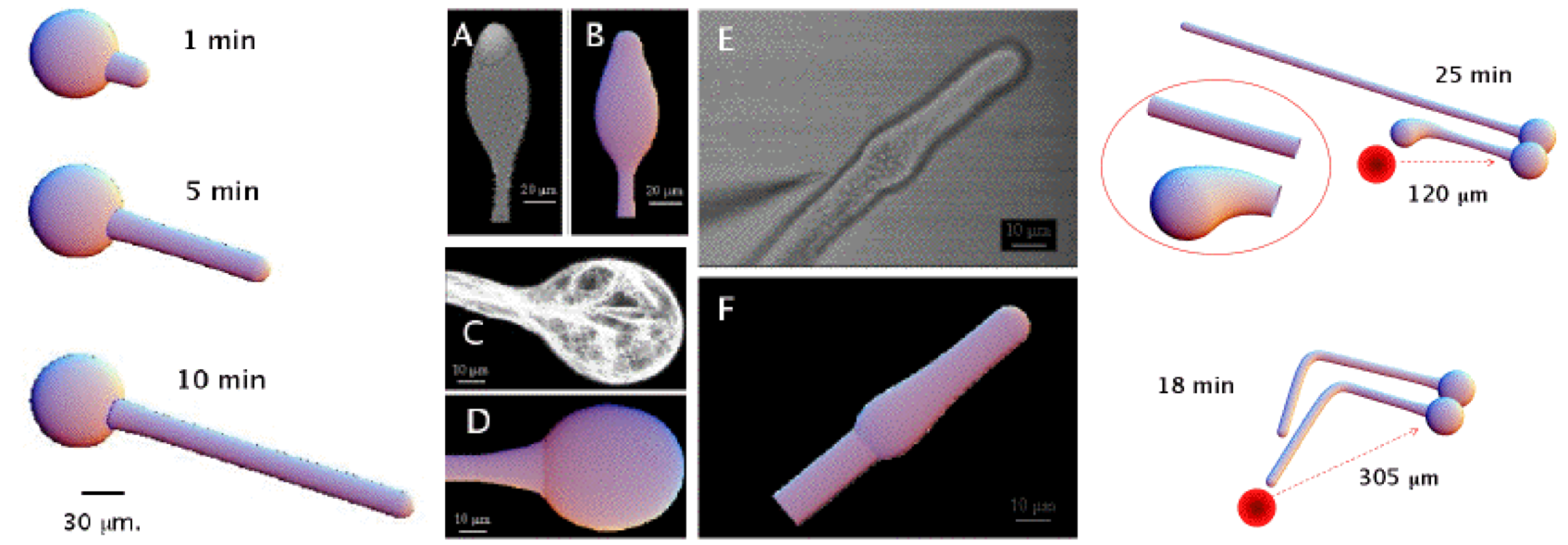
After getting his degree he remained in the group to close some loose-ends of his research project. Meanwhile, he was admitted to the PhD Program in Computational Biology in October 2005 and went on to do his Ph.D. with Stan Maree in the University of Utrecht. After spending some time, as a researcher at the Algarve University in Portugal, he joined the private sector, as senior data scientist.
 Tiago Carvalho did a four-month internship with me at the Instituto Gulbenkian de Ciência as part of his initial training in the Programa Gulbenkian de Doutoramento em Biomedicina. During this time, he modelled the stochastic monoallelic expression of cytokine genes. He went on to work on his thesis with Dean Buonomano at UCLA, studying the plasticity of neuronal input-output functions driven by changes in the balance between excitation and inhibition. Alongside his academic work, he was involved in extracurricular initiatives, including Papaformigas, a database of students from Portuguese PhD programmes. After completing his PhD, he founded and is now CEO of LabOrders. Tiago remains, to me, a prime example of resourcefulness and entrepreneurship.
Tiago Carvalho did a four-month internship with me at the Instituto Gulbenkian de Ciência as part of his initial training in the Programa Gulbenkian de Doutoramento em Biomedicina. During this time, he modelled the stochastic monoallelic expression of cytokine genes. He went on to work on his thesis with Dean Buonomano at UCLA, studying the plasticity of neuronal input-output functions driven by changes in the balance between excitation and inhibition. Alongside his academic work, he was involved in extracurricular initiatives, including Papaformigas, a database of students from Portuguese PhD programmes. After completing his PhD, he founded and is now CEO of LabOrders. Tiago remains, to me, a prime example of resourcefulness and entrepreneurship.
 When Rui Gardner returned to Portugal after conducting his PhD research with Armindo Salvador and Michael Savageau in Michigan, I invited him to make use of the material and intellectual resources of my group while writting his thesis. After successfully defending it, he stayed on as a postdoc for a couple of years. During this time, he made a risky yet farsighted move: he applied for the position of manager of the institute’s flow cytometry facility. In that role, he took the facility to a whole new level before moving on to the Memorial Sloan Kettering Cancer Center, where he is the Head of the Flow Cytometry Core Facility.
When Rui Gardner returned to Portugal after conducting his PhD research with Armindo Salvador and Michael Savageau in Michigan, I invited him to make use of the material and intellectual resources of my group while writting his thesis. After successfully defending it, he stayed on as a postdoc for a couple of years. During this time, he made a risky yet farsighted move: he applied for the position of manager of the institute’s flow cytometry facility. In that role, he took the facility to a whole new level before moving on to the Memorial Sloan Kettering Cancer Center, where he is the Head of the Flow Cytometry Core Facility.
 Nuno Sepúlveda, who trained as statistician, joined my group in 2001 as a research technician and soon became involved in a range of modeling projects, from AIDS pathogenesis to antigen-receptor gene recombination in lymphocyte precursors. While contributing to these projects, he pursued his Master’s in Applied Mathematics (Statistics and Applications), working on an allelic penetrance approach to genetic diseases, under the supervision of Carlos Penha-Gonçalves (IGC) and Daniel Paulino (IST). Motivated by his growing interest in immunology, Nuno embarked on a PhD project under my supervision, aiming to unify immunophysiology and genetics within a shared mathematical framework to clarify causal links between genetic variation and autoimmune disease. He earned his PhD from the University of Porto in 2019 with his thesis entitled "How is the T-cell repertoire shaped?" After a short period in the private sector, he returned to academia as a postdoctoral researcher at the London School of Hygiene and Tropical Medicine, where he worked for several years. He is currently a professor at the University of Warsaw, Poland.
Nuno Sepúlveda, who trained as statistician, joined my group in 2001 as a research technician and soon became involved in a range of modeling projects, from AIDS pathogenesis to antigen-receptor gene recombination in lymphocyte precursors. While contributing to these projects, he pursued his Master’s in Applied Mathematics (Statistics and Applications), working on an allelic penetrance approach to genetic diseases, under the supervision of Carlos Penha-Gonçalves (IGC) and Daniel Paulino (IST). Motivated by his growing interest in immunology, Nuno embarked on a PhD project under my supervision, aiming to unify immunophysiology and genetics within a shared mathematical framework to clarify causal links between genetic variation and autoimmune disease. He earned his PhD from the University of Porto in 2019 with his thesis entitled "How is the T-cell repertoire shaped?" After a short period in the private sector, he returned to academia as a postdoctoral researcher at the London School of Hygiene and Tropical Medicine, where he worked for several years. He is currently a professor at the University of Warsaw, Poland.
Nuno has long been my statistics guru. When asked to teach statistics courses for PhD programs at the Instituto Gulbenkian de Ciência (in Oeiras and Cidade da Praia) I often invited Nuno to join me. Our close collaboration and mutual trust, built over years of shared research and teaching, allowed us to step in for one another mid-lecture without missing a beat.
 Tiago Paixão joined my group at the Instituto Gulbenkian de Ciência as a PhD student, in 2023. He arrived with a solid education in physics and dynamical systems, having a degree in Physical Engineering from the IST and an internship at Rui Dilão's Non-linear Dynamics Group at IST, where he studied the geometry of the Lorenz attractor. In my group, he investigated the origins, mechanisms, and potential adaptive value of non-genetic heterogeneity in isogenic cell populations. Challenging the prevailing genetic reductionism in molecular biology, he explored how stochastic fluctuations in molecular interactions—particularly those involving low-copy-number species like chromosomes—lead to substantial phenotypic variation even among genetically identical cells in uniform environments. His work examined the phenomenon of pseudo monoallelic expression, especially in cytokine genes, where expression from individual alleles varies stochastically and lacks heritable stability. Tiago demonstrated that existing models of stochastic gene expression failed to account for these patterns and proposed a novel transcription model incorporating chromatin dynamics and allele independence that better fit experimental observations. He also addressed the empirical finding that protein copy numbers in single cells follow a lognormal distribution, a result incompatible with simple Poissonian models. Through mathematical modeling of multistep cellular processes, he showed that multiplicative noise inherent in these pathways naturally gives rise to lognormal distributions. Tiago further analyzed how variability in protein levels affects the behavior of signaling modules, demonstrating that population-level averages can obscure critical aspects of single-cell dynamics, leading to flawed mechanistic inferences. Finally, he explored whether such heterogeneity confers a selective advantage, showing that higher variance in protein copy numbers can accelerate somatic adaptation to environmental changes. His findings suggest that the structure of intracellular noise may itself be subject to evolutionary tuning, offering new perspectives on the interplay between stochasticity, cellular function, and adaptation.Tiago was awarded his doctoral degree in Biomedicine from the University of Porto in 2007 for his thesis entitled: "The Stochastic Basis of Somatic Variation". The thesis was short listed and received a Honourable Mention in the Reinhart Heinrich Award for the best doctoral thesis in any area of mathematical or theoretical Biology of the European Society for Mathematical and Theoretical Biology.
Tiago Paixão joined my group at the Instituto Gulbenkian de Ciência as a PhD student, in 2023. He arrived with a solid education in physics and dynamical systems, having a degree in Physical Engineering from the IST and an internship at Rui Dilão's Non-linear Dynamics Group at IST, where he studied the geometry of the Lorenz attractor. In my group, he investigated the origins, mechanisms, and potential adaptive value of non-genetic heterogeneity in isogenic cell populations. Challenging the prevailing genetic reductionism in molecular biology, he explored how stochastic fluctuations in molecular interactions—particularly those involving low-copy-number species like chromosomes—lead to substantial phenotypic variation even among genetically identical cells in uniform environments. His work examined the phenomenon of pseudo monoallelic expression, especially in cytokine genes, where expression from individual alleles varies stochastically and lacks heritable stability. Tiago demonstrated that existing models of stochastic gene expression failed to account for these patterns and proposed a novel transcription model incorporating chromatin dynamics and allele independence that better fit experimental observations. He also addressed the empirical finding that protein copy numbers in single cells follow a lognormal distribution, a result incompatible with simple Poissonian models. Through mathematical modeling of multistep cellular processes, he showed that multiplicative noise inherent in these pathways naturally gives rise to lognormal distributions. Tiago further analyzed how variability in protein levels affects the behavior of signaling modules, demonstrating that population-level averages can obscure critical aspects of single-cell dynamics, leading to flawed mechanistic inferences. Finally, he explored whether such heterogeneity confers a selective advantage, showing that higher variance in protein copy numbers can accelerate somatic adaptation to environmental changes. His findings suggest that the structure of intracellular noise may itself be subject to evolutionary tuning, offering new perspectives on the interplay between stochasticity, cellular function, and adaptation.Tiago was awarded his doctoral degree in Biomedicine from the University of Porto in 2007 for his thesis entitled: "The Stochastic Basis of Somatic Variation". The thesis was short listed and received a Honourable Mention in the Reinhart Heinrich Award for the best doctoral thesis in any area of mathematical or theoretical Biology of the European Society for Mathematical and Theoretical Biology.
After an international postdoctoral career he is back to Portugal, where he heads the Quantitative Biology Unit at the GIMM.
 Back in early 2000's, Dejan Milutinović was a PhD student of the Instituto Superior Técnico with Michael Athans and Pedro Lima. During is research we collaborated in the development of models of the dynamics of antigen receptors in T cell populations that served as inspiration to develop control strategies for distributed robotic systems. Dejan's' PhD dissertation was voted for the first runner-up for the best PhD thesis of European Robotics in 2004 by EURON, the network of over 150 European robotics research groups. Dejan is a professor in the Department of Electrical and Computer Engineering, UC Santa Cruz. You can find him here.
Back in early 2000's, Dejan Milutinović was a PhD student of the Instituto Superior Técnico with Michael Athans and Pedro Lima. During is research we collaborated in the development of models of the dynamics of antigen receptors in T cell populations that served as inspiration to develop control strategies for distributed robotic systems. Dejan's' PhD dissertation was voted for the first runner-up for the best PhD thesis of European Robotics in 2004 by EURON, the network of over 150 European robotics research groups. Dejan is a professor in the Department of Electrical and Computer Engineering, UC Santa Cruz. You can find him here.
 Andreia Lino joined my group as an undergraduate student, from September 2001 to July 2002, to conduct her final-year research project under the joint supervision of Matthias Haury and me. Her goal was to quantify key aspects of T-lymphocyte activation using an integrated approach that combined experimental bench work with mathematical modelling. She quantitatively characterised the heterogeneity and kinetics of peptide loading onto MHC class I molecules by antigen-presenting cells and mathematically modelled the kinetics of T-cell receptor downregulation in response to different ligands. Her final report, entitled "Estudo do mecanismo de activação dos linfócitos T", was graded among the highest in her cohort.
Andreia received her Biochemistry degree in 2002 and went on to pursue a PhD with Juan Lafaille at the New York University Medical Center. After completing her doctorate, she returned to the Instituto Gulbenkian de Ciência as a postdoctoral fellow in Jocelyne Demengeot’s lab. She is currently a Postdoctoral Fellow at the Deutsches Rheuma-Forschungszentrum.
Andreia Lino joined my group as an undergraduate student, from September 2001 to July 2002, to conduct her final-year research project under the joint supervision of Matthias Haury and me. Her goal was to quantify key aspects of T-lymphocyte activation using an integrated approach that combined experimental bench work with mathematical modelling. She quantitatively characterised the heterogeneity and kinetics of peptide loading onto MHC class I molecules by antigen-presenting cells and mathematically modelled the kinetics of T-cell receptor downregulation in response to different ligands. Her final report, entitled "Estudo do mecanismo de activação dos linfócitos T", was graded among the highest in her cohort.
Andreia received her Biochemistry degree in 2002 and went on to pursue a PhD with Juan Lafaille at the New York University Medical Center. After completing her doctorate, she returned to the Instituto Gulbenkian de Ciência as a postdoctoral fellow in Jocelyne Demengeot’s lab. She is currently a Postdoctoral Fellow at the Deutsches Rheuma-Forschungszentrum.
 Vanessa Oliveira was a researcher in my group from April 1998 to March 1999. She held a degree in Biochemistry from the Faculty of Sciences of the University of Lisbon. Her project aimed to experimentally test our mathematical models of regulatory T cell dynamics and interactions, in collaboration with Jocelyne Demengeot’s group. Vanessa quantitatively analysed several aspects of in vitro interactions between regulatory T cells, conventional T cells, and antigen-presenting cells. Her most significant contribution was the experimental validation of the Crossregulation model, which proposed that conventional T cells act as growth factors for regulatory T cells. This validation was a key stepping stone for subsequent theoretical developments in the group.
Vanessa Oliveira was a researcher in my group from April 1998 to March 1999. She held a degree in Biochemistry from the Faculty of Sciences of the University of Lisbon. Her project aimed to experimentally test our mathematical models of regulatory T cell dynamics and interactions, in collaboration with Jocelyne Demengeot’s group. Vanessa quantitatively analysed several aspects of in vitro interactions between regulatory T cells, conventional T cells, and antigen-presenting cells. Her most significant contribution was the experimental validation of the Crossregulation model, which proposed that conventional T cells act as growth factors for regulatory T cells. This validation was a key stepping stone for subsequent theoretical developments in the group.
Vanessa later pursued a PhD at Oxford University with Kathryn Wood, focusing on the role of regulatory T cells in transplantation tolerance. She returned to Portugal as a postdoctoral fellow in Luís Graça’s lab at the Institute for Molecular Medicine in Lisbon. Meanwhile, she earned a degree in Medicine from the University of Lisbon and is now a practising medical doctor.
 Kalet Leon got a degree in Physic from the University of Havana in Cuba in 1997, and joined the Center for Molecular Immunology as young researcher. We met in person during an international workshop on Immunotherapy at the CIM, and he joined my lab in part-time as a Ph.D. student. The aim of his project was to understand the mechanism of tolerance mediated by regulatory T cells. To achieve this goal he had to develop a model for the interactions in multicellular conjugates. He demonstrated on formal grounds that regulatory cells grow as a function of the cells they suppress from inducing autoimmunity. He simulated the repertoire selection of regulatory T cells showing that in the thymus regulatory cells could be produced randomly, independently of the specificity, without compromising self-nonself discrimination. Finally, he has shown that the crosstalk mechanism between regulatory T cells and pathogenic T cells naturally predicts the existence of an inverse correlation between infections and autoimmune diseases. He got a Ph.D. in Biomedicine from the University of Porto in 2002 for his thesis entitled "A quantitative approach to dominant tolerance". After that, he continued as a postdoc in the group, still in part-time, because he was assigned a new job at the CIM back in Cuba. He is now the Deputy Director of the CIM.
Kalet Leon got a degree in Physic from the University of Havana in Cuba in 1997, and joined the Center for Molecular Immunology as young researcher. We met in person during an international workshop on Immunotherapy at the CIM, and he joined my lab in part-time as a Ph.D. student. The aim of his project was to understand the mechanism of tolerance mediated by regulatory T cells. To achieve this goal he had to develop a model for the interactions in multicellular conjugates. He demonstrated on formal grounds that regulatory cells grow as a function of the cells they suppress from inducing autoimmunity. He simulated the repertoire selection of regulatory T cells showing that in the thymus regulatory cells could be produced randomly, independently of the specificity, without compromising self-nonself discrimination. Finally, he has shown that the crosstalk mechanism between regulatory T cells and pathogenic T cells naturally predicts the existence of an inverse correlation between infections and autoimmune diseases. He got a Ph.D. in Biomedicine from the University of Porto in 2002 for his thesis entitled "A quantitative approach to dominant tolerance". After that, he continued as a postdoc in the group, still in part-time, because he was assigned a new job at the CIM back in Cuba. He is now the Deputy Director of the CIM.
 João Sousa joined me at the Instituto Gulbenkian de Ciência in April 1998 as my very first PhD student. His project aimed to understand how antigen and cytokine receptor signalling shape T cell activation, differentiation, and population dynamics. To that end, João developed the first cell biology modelling framework that integrated signal transduction, intercellular interactions, and cell population dynamics. João modelled the interference between the responses to the cytokines IL-2 and IL-4, whose receptors share a common chain, showing that this interference facilitates plasticity of immune responses and memory. He used accurate data on TCR downregulation to unravel the quantitative details of the kinetics of TCR triggering and downregulation. Based on this quantitative studies he proposed a hypothesis according to which the T cell uses multiple-TCR triggers to measure different properties of the antigen. Finally, he put forward a model for the T cell response to serial conjugation with the antigen presenting cell. João's thesis became the steping stone of several research lines at the Group. He received his PhD in Theoretical Biochemistry from the University of Lisbon in October 2003, with a thesis entitled "Modeling the antigen and cytokine receptors signalling processes and their propagation to lymphocyte population dynamics." In the group, João also served as our system administrator, and after graduating he becaming increasingly involved with the IT Unit at IGC eventually becoming its leader. He his now an IT consultant providing specialised advise to research institutions.
João Sousa joined me at the Instituto Gulbenkian de Ciência in April 1998 as my very first PhD student. His project aimed to understand how antigen and cytokine receptor signalling shape T cell activation, differentiation, and population dynamics. To that end, João developed the first cell biology modelling framework that integrated signal transduction, intercellular interactions, and cell population dynamics. João modelled the interference between the responses to the cytokines IL-2 and IL-4, whose receptors share a common chain, showing that this interference facilitates plasticity of immune responses and memory. He used accurate data on TCR downregulation to unravel the quantitative details of the kinetics of TCR triggering and downregulation. Based on this quantitative studies he proposed a hypothesis according to which the T cell uses multiple-TCR triggers to measure different properties of the antigen. Finally, he put forward a model for the T cell response to serial conjugation with the antigen presenting cell. João's thesis became the steping stone of several research lines at the Group. He received his PhD in Theoretical Biochemistry from the University of Lisbon in October 2003, with a thesis entitled "Modeling the antigen and cytokine receptors signalling processes and their propagation to lymphocyte population dynamics." In the group, João also served as our system administrator, and after graduating he becaming increasingly involved with the IT Unit at IGC eventually becoming its leader. He his now an IT consultant providing specialised advise to research institutions.
 João Sousa and me, with Rute and other off camera colleagues, in the summer of 1999.
João Sousa and me, with Rute and other off camera colleagues, in the summer of 1999.
 António Coutinho, the thought-provoking immunologist, the compelling lecturer, the welcoming host, the scientist with an intellectual tour du monde, the Mentor, the visionary and charismatic Director of the IGC, the constructive yet uncompromising advisor, the friend, is to big for a small paragraph. As a PhD student squatting in his lab at the Unité d’Immunologie de l’ Institut Pasteur or as an independent scientist at the Instituto Gulbenkian de Ciência, I have been enjoying the privilege of many, many dialogues—in the absence of a better word for “conversas”—with António that invariably started with: “do you have five minutes?”.
António Coutinho, the thought-provoking immunologist, the compelling lecturer, the welcoming host, the scientist with an intellectual tour du monde, the Mentor, the visionary and charismatic Director of the IGC, the constructive yet uncompromising advisor, the friend, is to big for a small paragraph. As a PhD student squatting in his lab at the Unité d’Immunologie de l’ Institut Pasteur or as an independent scientist at the Instituto Gulbenkian de Ciência, I have been enjoying the privilege of many, many dialogues—in the absence of a better word for “conversas”—with António that invariably started with: “do you have five minutes?”.
 José Faro played a major role in my research, being a lifelong colleague and friend. I first met him when he visited Arala Chaves’ Laboratory of Immunology, where he gave an informal seminar on immune system modelling. This was my first exposure to theoretical immunology. We reconnected a couple of years later at a meeting in Luminy, by which time I was already working on my thesis at the Pasteur Institute. We applied for a travel grant that supported our collaboration, allowing him to visit me in Paris and me to visit him at the Physics Department of the University of Salamanca. We worked on topics adjacent to my thesis and coauthored a few papers. The time spent working and discussing with José was formative: he challenged me to explore new areas in immunology and modelling, and helped me to be more critical about my work. When I started my own group at the Instituto Gulbenkian de Ciência, I invited José to join us so that my students could benefit from the kind of mentorship he had given me. We were fortunate that he accepted to be a recurrent visiting scientist from 1998 until 2003, when he went on to establish his own Systems Immunology group at the institute.
José Faro played a major role in my research, being a lifelong colleague and friend. I first met him when he visited Arala Chaves’ Laboratory of Immunology, where he gave an informal seminar on immune system modelling. This was my first exposure to theoretical immunology. We reconnected a couple of years later at a meeting in Luminy, by which time I was already working on my thesis at the Pasteur Institute. We applied for a travel grant that supported our collaboration, allowing him to visit me in Paris and me to visit him at the Physics Department of the University of Salamanca. We worked on topics adjacent to my thesis and coauthored a few papers. The time spent working and discussing with José was formative: he challenged me to explore new areas in immunology and modelling, and helped me to be more critical about my work. When I started my own group at the Instituto Gulbenkian de Ciência, I invited José to join us so that my students could benefit from the kind of mentorship he had given me. We were fortunate that he accepted to be a recurrent visiting scientist from 1998 until 2003, when he went on to establish his own Systems Immunology group at the institute.
By the end of my PhD, Caroline Brissac became my colleague at the Pasteur Institute, as John Stewart's other student. We overlapped for a couple of years in Paris, and she later visited me in Utrecht and eventually in Oeiras, after I established myself at Instituto Gulbenkian de Ciência. It is fair to say that Caroline was my first experience in (co-)supervising a research student; an experience which, as we all know, goes beyond scientific guidance and involves the abilities to listen, motivate, and support the other person. She did excellent work on estimating antibody repertoire diversity using the Panama blot, an early immunoproteomics method developed at Pasteur by Matthias Haury and Alberto Nobrega.
 John Stewart was my PhD supervisor and mentor at the Unité d'Immunobiologie at the Pasteur Institute. One of the most intelligent people I’ve ever met, he was also iconoclastic, unapologetically unconventional, and genuinely unafraid to speak his mind or to be proven wrong. I first encountered him at a meeting in Salamanca organized by Miguel Ángel Marcos. There, in his unmistakable style, he gave a provocative (and long, very long) talk on the evolution of the immune system, arguing boldly that the vertebrate immune system did not evolve to fight infection. He supported this claim with detailed insights into the evolutionary innovations required to produce a fully-fledged lymphocyte-based immune system.
John Stewart was my PhD supervisor and mentor at the Unité d'Immunobiologie at the Pasteur Institute. One of the most intelligent people I’ve ever met, he was also iconoclastic, unapologetically unconventional, and genuinely unafraid to speak his mind or to be proven wrong. I first encountered him at a meeting in Salamanca organized by Miguel Ángel Marcos. There, in his unmistakable style, he gave a provocative (and long, very long) talk on the evolution of the immune system, arguing boldly that the vertebrate immune system did not evolve to fight infection. He supported this claim with detailed insights into the evolutionary innovations required to produce a fully-fledged lymphocyte-based immune system.  By that time, I had already read his work on immune network modelling and was fascinated by the breadth of his interests. When I began my PhD, John gave me total freedom to shape my research. He visited the Pasteur Institute regularly, weekly at first, then less and less frequently. During each visit, I would walk him through my progress or give him a manuscript to work on. These sessions would quickly unfold (often derail) into long, intense discussions about whatever was on either of our minds at the time. On one of those visits, early on, I found John making photocopies of a publication and discovered, to my surprise, that he was the Stewart of the Elston-Stewart algorithm—an essential method in pedigree genetics. His freshly published paper had been commissioned by the journal Human Heredity to mark the 20th anniversary of that landmark publication. It turned out that John, who was trained in mathematical physics and genetics of kidney physiology in Cambridge, had walked away from statistical genetics, and from research altogether, at the height of his career, disillusioned with the direction the scientific enterprise was taking, driven by reductionist molecular biology. He spent many years outside academia before returning, drawn back by his interactions with Francisco Varela. His interest in immunology wasn’t driven by a desire to understand the immune system per se, but rather by the opportunity it offered to explore and develop the concept of autopoiesis. For John, the immune system was a rich and concrete arena in which to think about how biological systems bring forth their identity. John's long hiatus explains why I was his very first PhD student, and also why he cultivated my independence and allowed me to take full responsibility for my own project—an approach to supervision reminiscent of an earlier era. In the early 2000s, John joined the Instituto Gulbenkian de Ciência as a visiting scientist, alongside Mel Cohn, Rod Langman, José Faro, and Zvi Grossman. His expertise in statistical genetics proved invaluable to several groups working on disease genetics, including those led by Carlos Penha-Gonçalves and Astrid Vicente. As always, John and I spent long hours in deep, critical conversations about our work; discussions that continued to influence the way I think about science and society.
By that time, I had already read his work on immune network modelling and was fascinated by the breadth of his interests. When I began my PhD, John gave me total freedom to shape my research. He visited the Pasteur Institute regularly, weekly at first, then less and less frequently. During each visit, I would walk him through my progress or give him a manuscript to work on. These sessions would quickly unfold (often derail) into long, intense discussions about whatever was on either of our minds at the time. On one of those visits, early on, I found John making photocopies of a publication and discovered, to my surprise, that he was the Stewart of the Elston-Stewart algorithm—an essential method in pedigree genetics. His freshly published paper had been commissioned by the journal Human Heredity to mark the 20th anniversary of that landmark publication. It turned out that John, who was trained in mathematical physics and genetics of kidney physiology in Cambridge, had walked away from statistical genetics, and from research altogether, at the height of his career, disillusioned with the direction the scientific enterprise was taking, driven by reductionist molecular biology. He spent many years outside academia before returning, drawn back by his interactions with Francisco Varela. His interest in immunology wasn’t driven by a desire to understand the immune system per se, but rather by the opportunity it offered to explore and develop the concept of autopoiesis. For John, the immune system was a rich and concrete arena in which to think about how biological systems bring forth their identity. John's long hiatus explains why I was his very first PhD student, and also why he cultivated my independence and allowed me to take full responsibility for my own project—an approach to supervision reminiscent of an earlier era. In the early 2000s, John joined the Instituto Gulbenkian de Ciência as a visiting scientist, alongside Mel Cohn, Rod Langman, José Faro, and Zvi Grossman. His expertise in statistical genetics proved invaluable to several groups working on disease genetics, including those led by Carlos Penha-Gonçalves and Astrid Vicente. As always, John and I spent long hours in deep, critical conversations about our work; discussions that continued to influence the way I think about science and society.
 Manuel Vilanova was my direct mentor during my final research project for the Biochemistry degree, carried out in Arala Chaves' Laboratory of Immunology. I learned a great deal from Manuel, far beyond protein purification, animal experimentation, and flow cytometry. I have vivid memories of our flow cytometry sessions at the EPICS Coulter machine, which the lab shared with the Immunology Department of Hospital Geral de Santo António.
Since we only had access after the hospital’s daytime operations ended, those long after hours sessions contributed to consolidate our friendship though discussions about immunology, science, and life. Manuel played a pivotal role in shaping my path, incouraging me to persue a PhD in immune system modelling. More recently, with the help of several colleagues, Manuel and I submitted a proposal for an innovative Master’s in Integrative and Systems Immunology. This initiative
prompted a reconnection with my alma mater, the University of Porto.
Manuel Vilanova was my direct mentor during my final research project for the Biochemistry degree, carried out in Arala Chaves' Laboratory of Immunology. I learned a great deal from Manuel, far beyond protein purification, animal experimentation, and flow cytometry. I have vivid memories of our flow cytometry sessions at the EPICS Coulter machine, which the lab shared with the Immunology Department of Hospital Geral de Santo António.
Since we only had access after the hospital’s daytime operations ended, those long after hours sessions contributed to consolidate our friendship though discussions about immunology, science, and life. Manuel played a pivotal role in shaping my path, incouraging me to persue a PhD in immune system modelling. More recently, with the help of several colleagues, Manuel and I submitted a proposal for an innovative Master’s in Integrative and Systems Immunology. This initiative
prompted a reconnection with my alma mater, the University of Porto.
 I discovered immunology through Professor Arala Chaves, with whom I did my undergraduate research placement. Arala was irreverent and unconventional. He would shock us from the very first encounter by stating, with unshakable conviction, that there were no good immunology textbooks. He claimed they were too simplistic and irreparably outdated. The corollary was that one should discover immunology through primary scientific literature. A herculean task for someone just starting out in the field, but one that taught us to recognise uncertainty and to consolidate our knowledge by confronting hypotheses with data.
He was the first person to make me reflect on the idea that infectious agents can teach us a great deal about the regulation of the immune system. Arala showed, time and again, that some pathogens, in order to chronically infect a host, manipulate its immune system. Bacteria and viruses produce or induce the host to produce mitogens that stimulate all lymphocytes indiscriminately, thereby suppressing the responses of those that are specific to the pathogen.
As a metaphor, Arala used judo, the martial art in which a weaker fighter uses the strength of a stronger opponent to subdue them. He argued that vaccines should be directed against these pathogen-derived mitogens, rather than against structural components, as is more conventional. This idea was so far removed from the textbook vision that we came to realise we shouldn't fear having original ideas. He also forced us to think about the immune response in a complex and non-linear way, which, in my case, led to a lasting interest in the mathematical modelling of the immune system and its regulation.
I discovered immunology through Professor Arala Chaves, with whom I did my undergraduate research placement. Arala was irreverent and unconventional. He would shock us from the very first encounter by stating, with unshakable conviction, that there were no good immunology textbooks. He claimed they were too simplistic and irreparably outdated. The corollary was that one should discover immunology through primary scientific literature. A herculean task for someone just starting out in the field, but one that taught us to recognise uncertainty and to consolidate our knowledge by confronting hypotheses with data.
He was the first person to make me reflect on the idea that infectious agents can teach us a great deal about the regulation of the immune system. Arala showed, time and again, that some pathogens, in order to chronically infect a host, manipulate its immune system. Bacteria and viruses produce or induce the host to produce mitogens that stimulate all lymphocytes indiscriminately, thereby suppressing the responses of those that are specific to the pathogen.
As a metaphor, Arala used judo, the martial art in which a weaker fighter uses the strength of a stronger opponent to subdue them. He argued that vaccines should be directed against these pathogen-derived mitogens, rather than against structural components, as is more conventional. This idea was so far removed from the textbook vision that we came to realise we shouldn't fear having original ideas. He also forced us to think about the immune response in a complex and non-linear way, which, in my case, led to a lasting interest in the mathematical modelling of the immune system and its regulation.